CDSeq – A novel complete deconvolution method for dissecting heterogeneous samples using gene expression data | RNA-Seq Blog
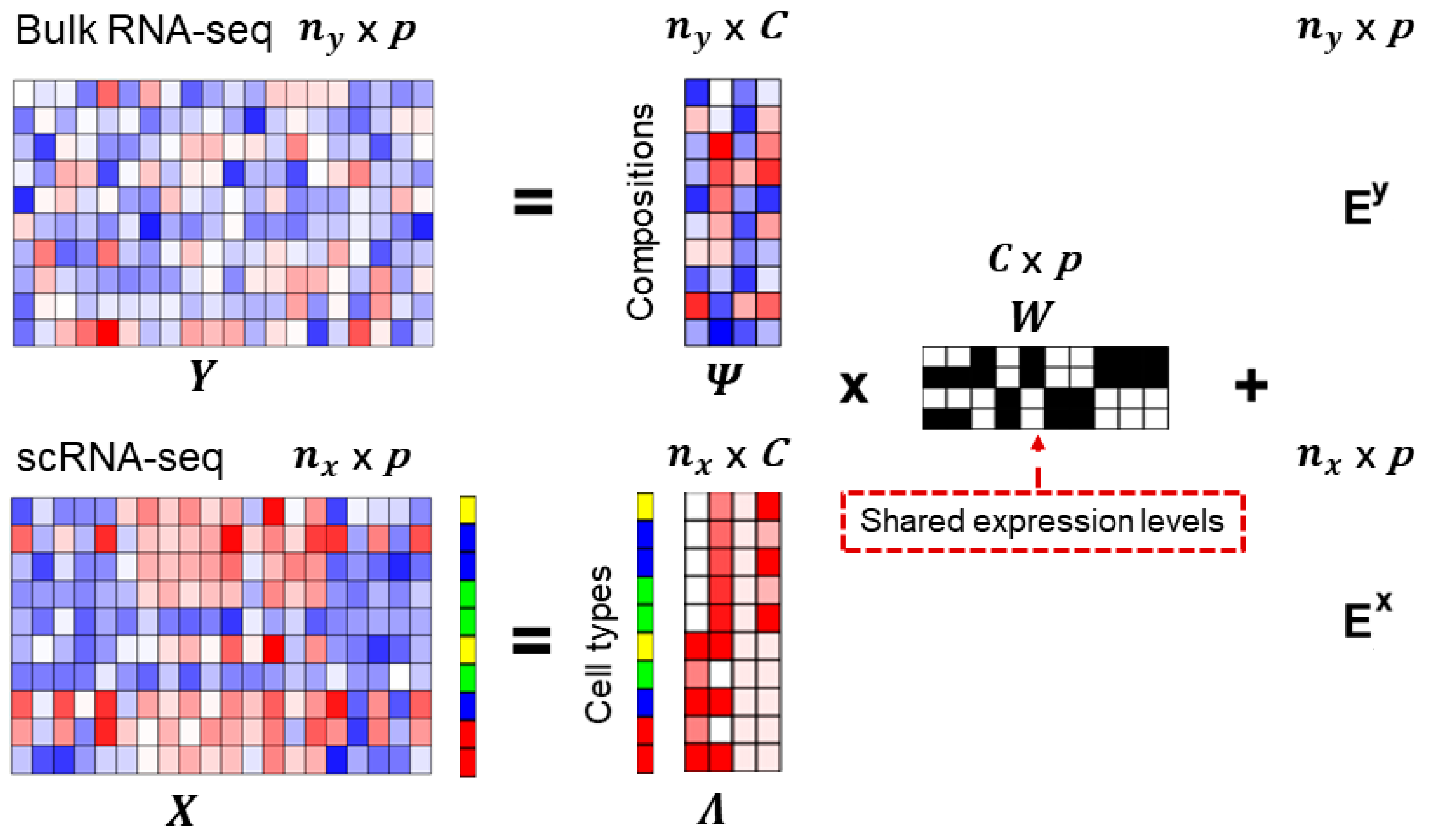
Cells | Free Full-Text | An Efficient and Flexible Method for Deconvoluting Bulk RNA-Seq Data with Single-Cell RNA-Seq Data

Integrative single-cell and bulk RNA-seq analysis in human retina identified cell type-specific composition and gene expression changes for age-related macular degeneration | bioRxiv
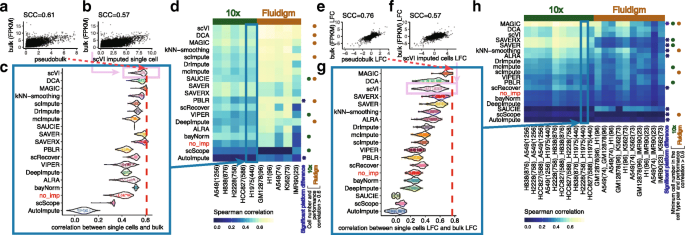
A systematic evaluation of single-cell RNA-sequencing imputation methods | Genome Biology | Full Text
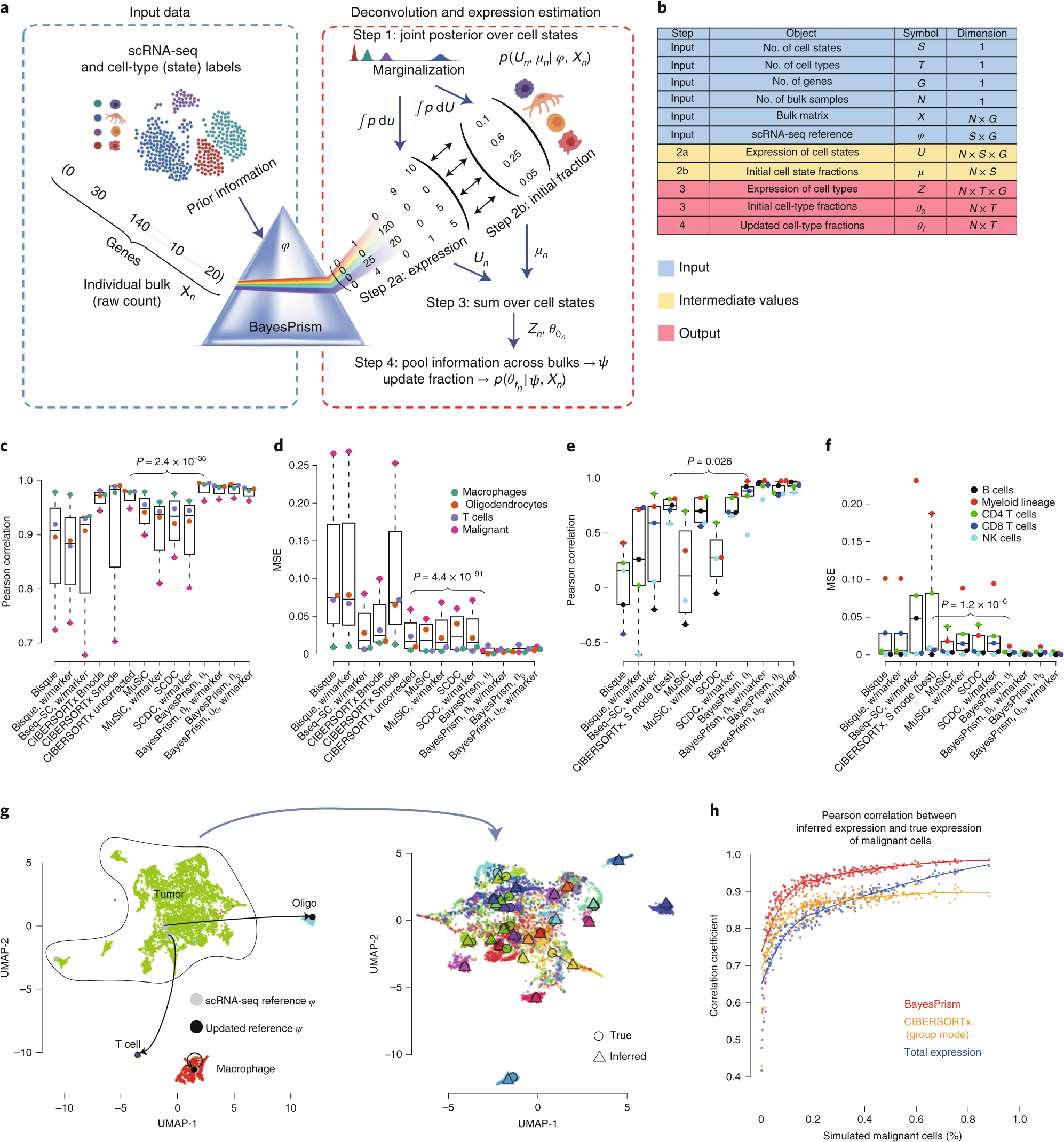
Cell type and gene expression deconvolution with BayesPrism enables Bayesian integrative analysis across bulk and single-cell RNA sequencing in oncology | Nature Cancer
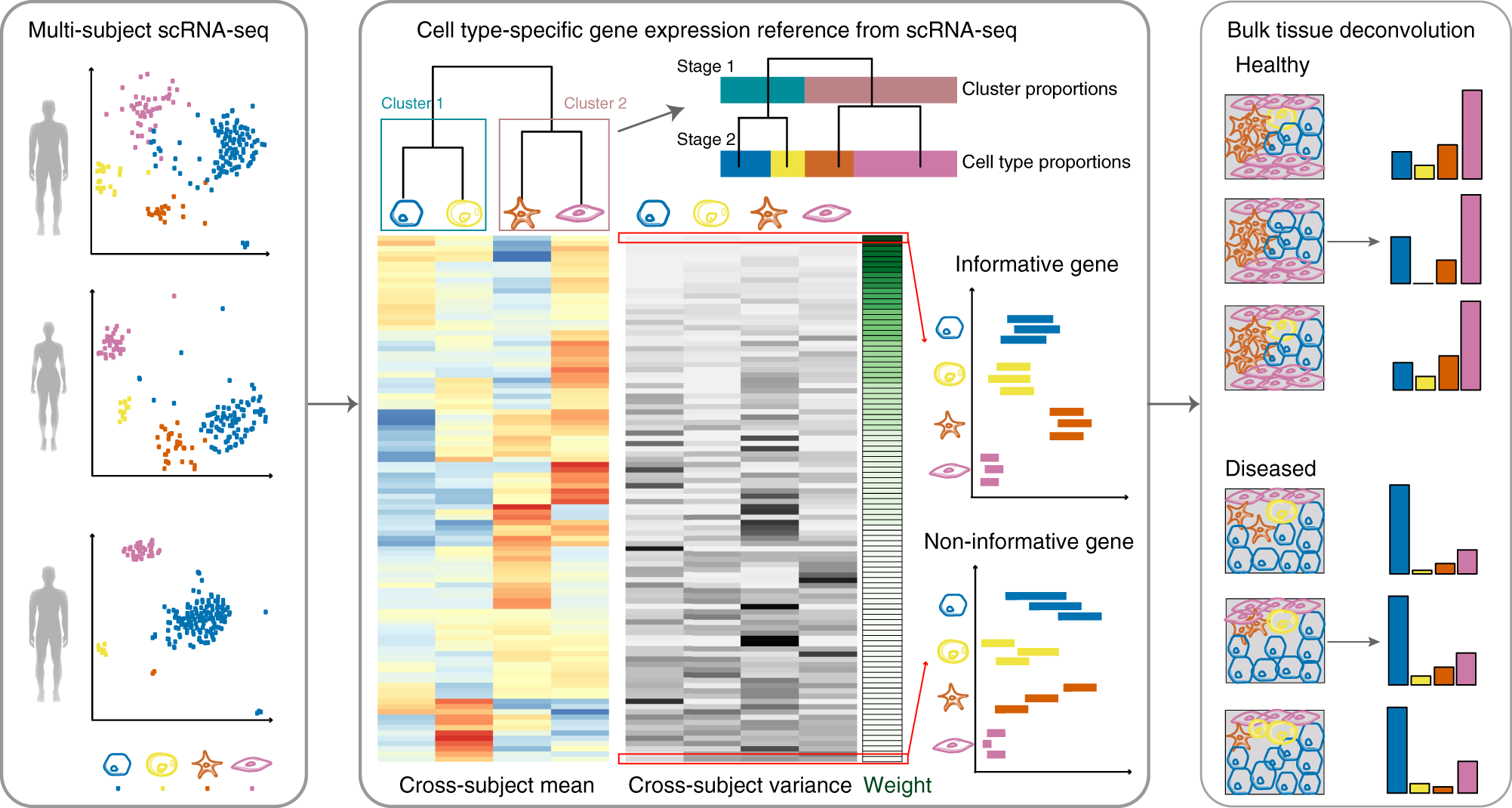
Bulk tissue cell type deconvolution with multi-subject single-cell expression reference | Nature Communications
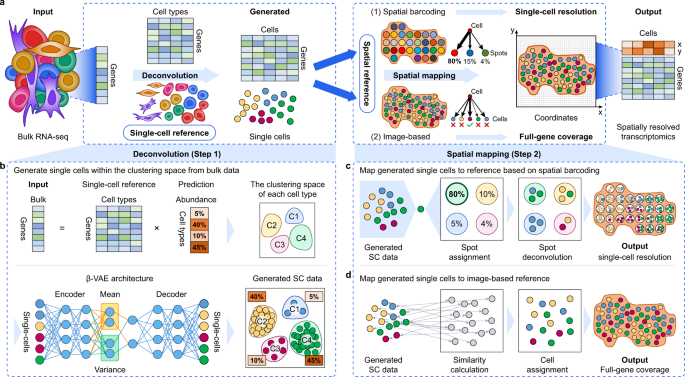
De novo analysis of bulk RNA-seq data at spatially resolved single-cell resolution | Nature Communications

SCDC – bulk gene expression deconvolution by multiple single-cell RNA sequencing references | RNA-Seq Blog
Detecting cell-type-specific allelic expression imbalance by integrative analysis of bulk and single-cell RNA sequencing data | PLOS Genetics