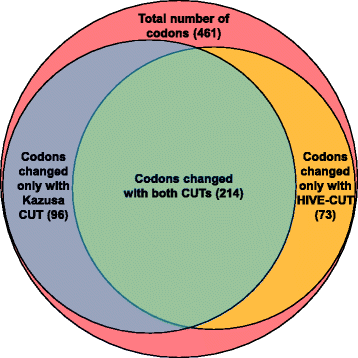
Codon Usage Optimization in the Prokaryotic Tree of Life: How Synonymous Codons Are Differentially Selected in Sequence Domains with Different Expression Levels and Degrees of Conservation | mBio
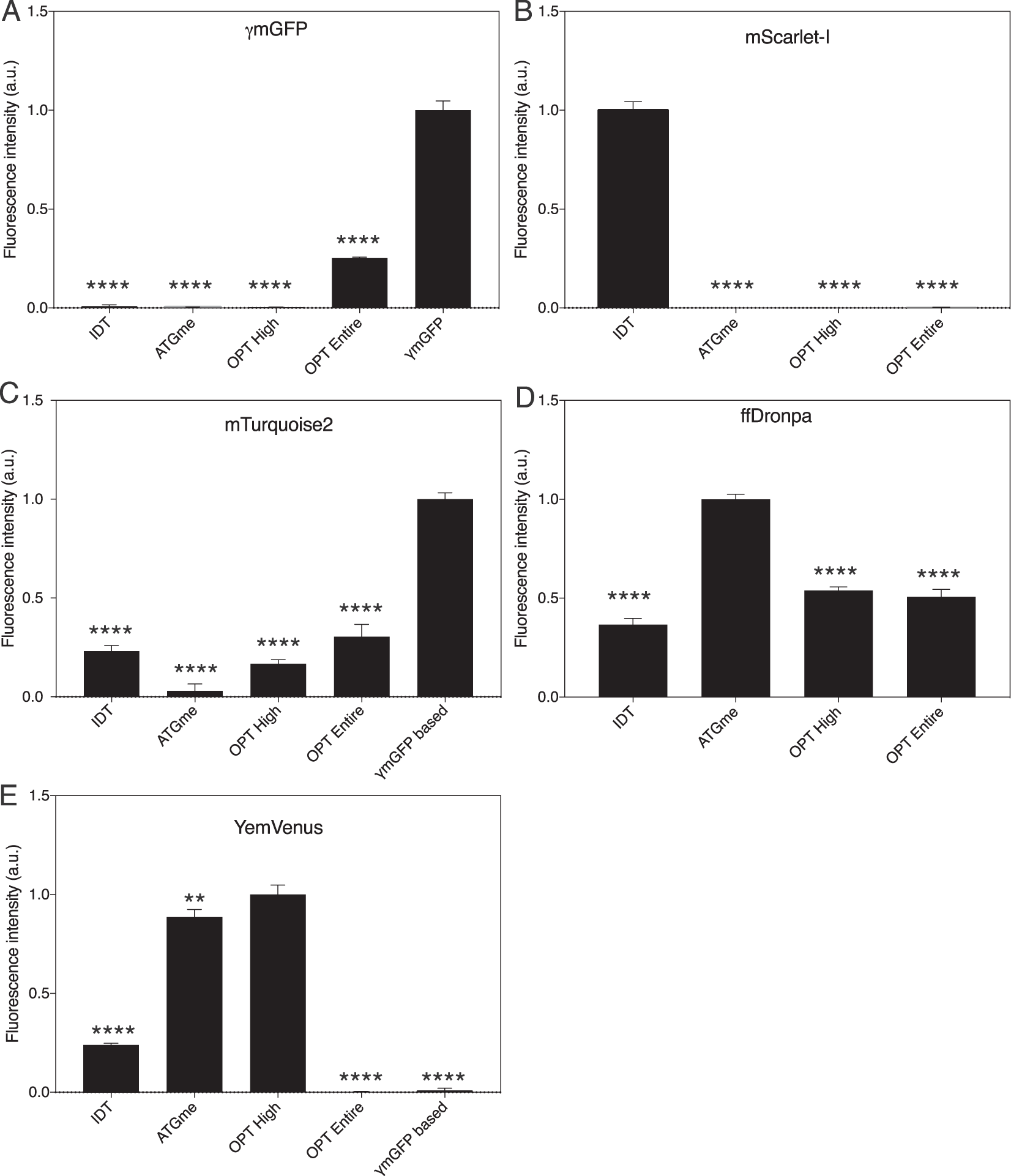
Presenting a codon-optimized palette of fluorescent proteins for use in Candida albicans | Scientific Reports
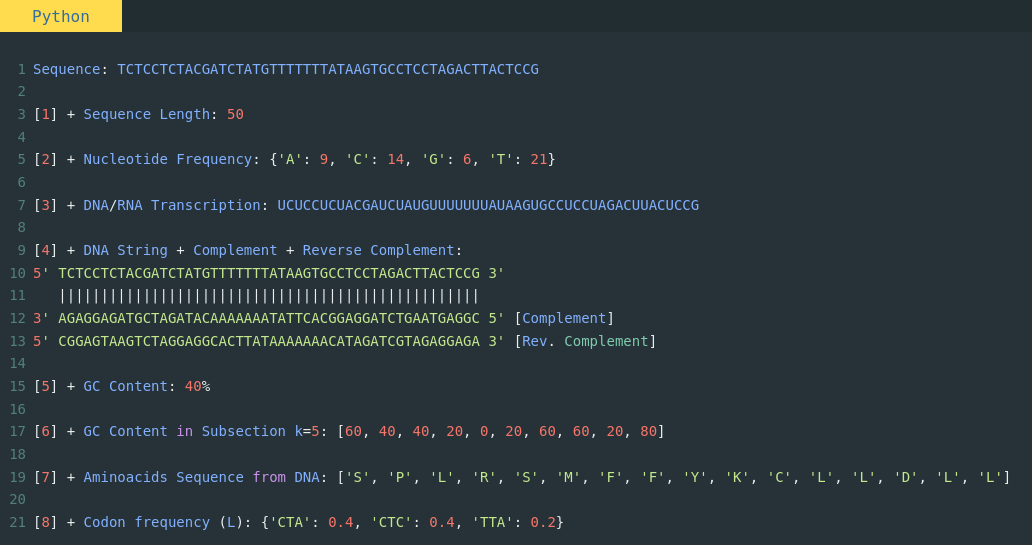
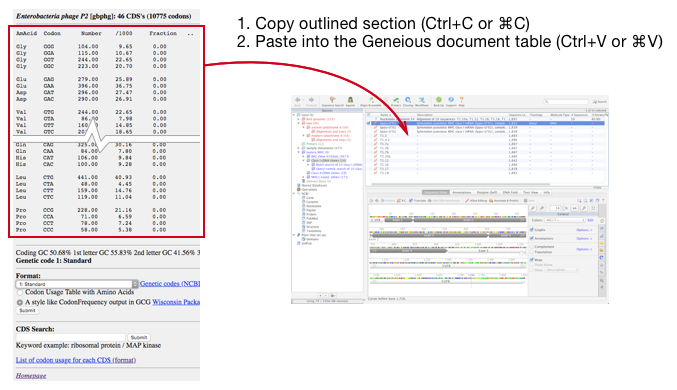
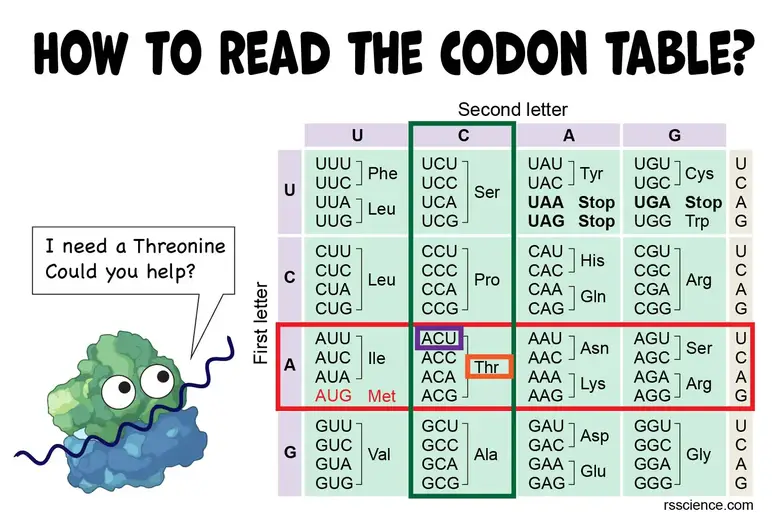
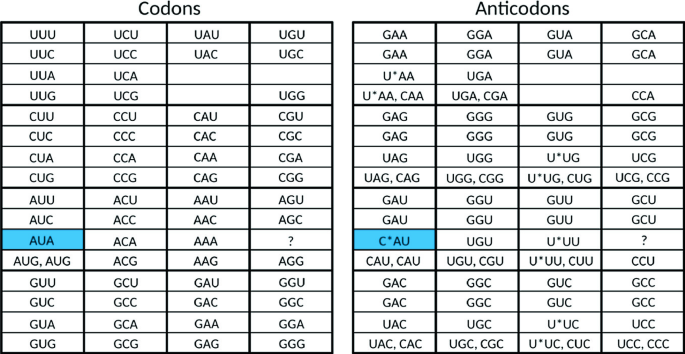
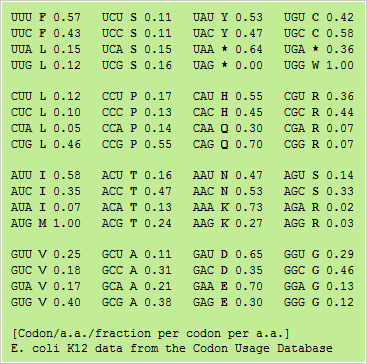
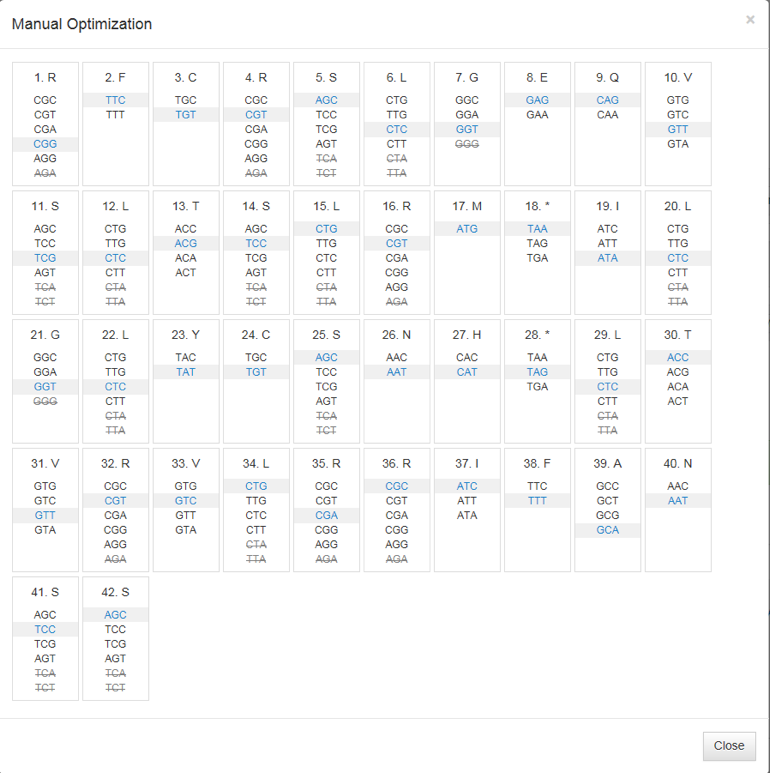
Bioinformatics in Python; DNA Toolkit. Part 4: Translation, Codon Usage | by rebelCoder | Python in Plain English

Comparison of codon usage. 1) Codons with low frequency ( , 10%) are... | Download Scientific Diagram

MinMax: A versatile tool for calculating and comparing synonymous codon usage and its impact on protein folding - Rodriguez - 2018 - Protein Science - Wiley Online Library
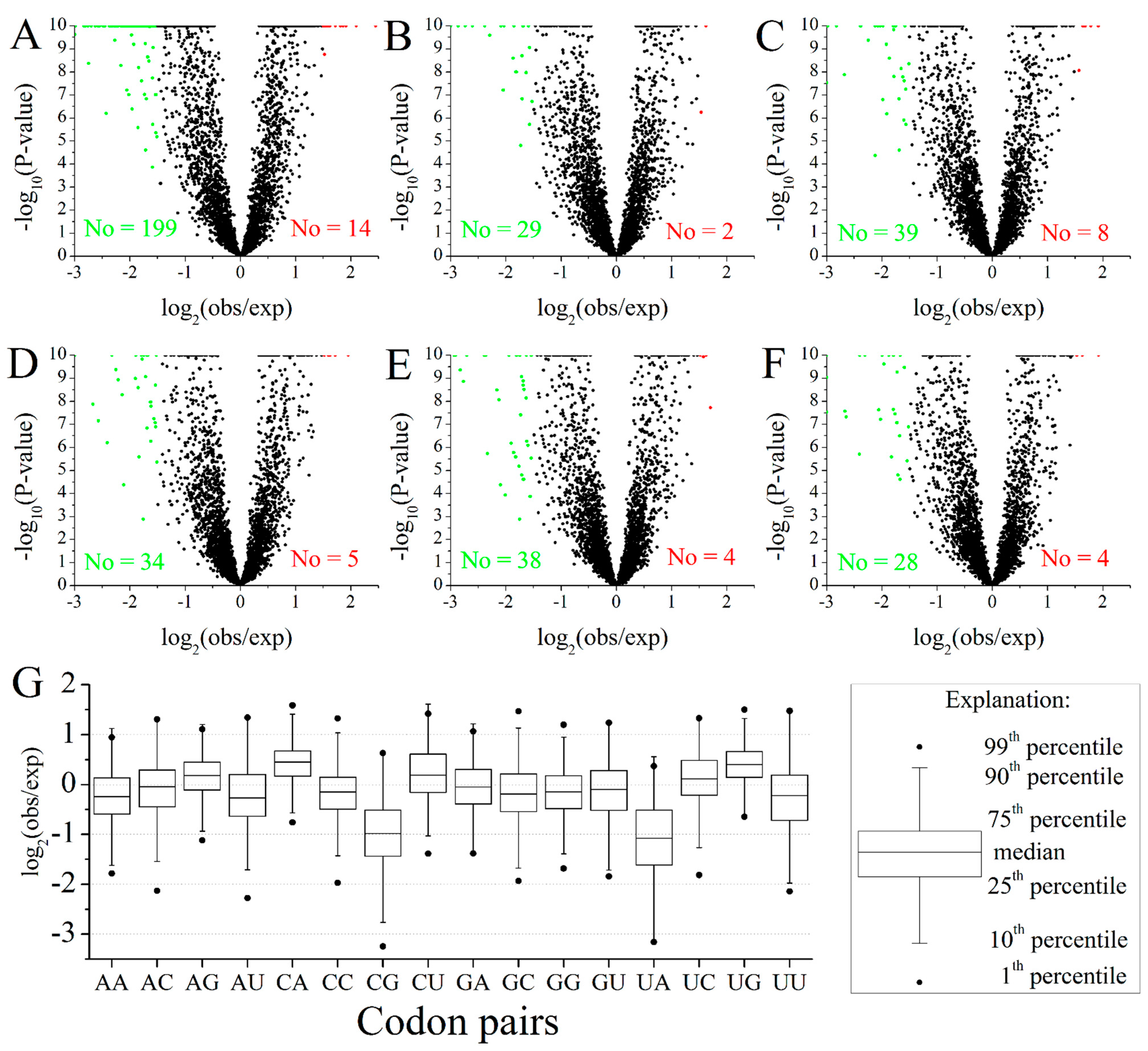
IJMS | Free Full-Text | A Comprehensive Analysis of Codon Usage Patterns in Blunt Snout Bream (Megalobrama amblycephala) Based on RNA-Seq Data
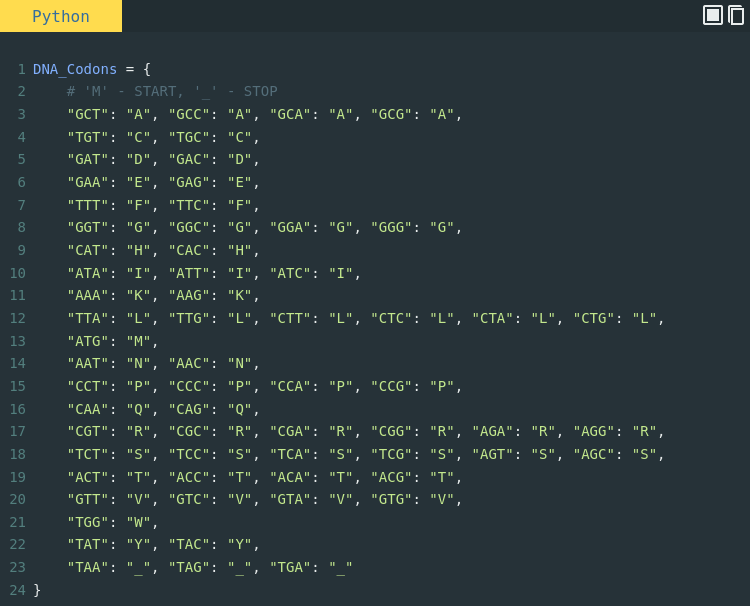
Bioinformatics in Python; DNA Toolkit. Part 4: Translation, Codon Usage | by rebelCoder | Python in Plain English


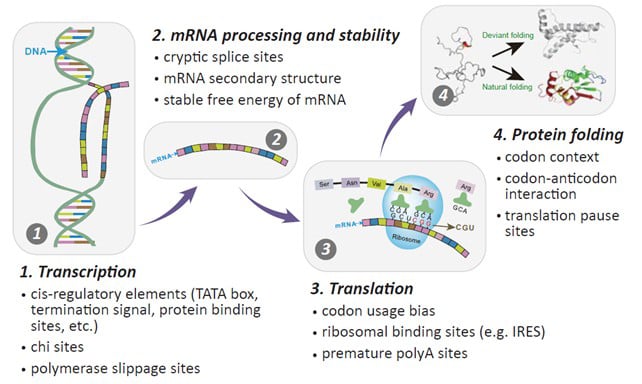
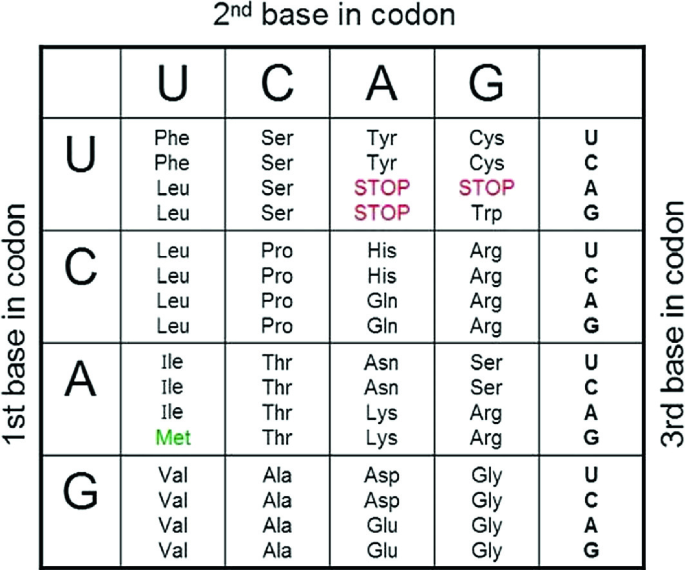




![Analysis of synonymous codon usage patterns in sixty-four different bivalve species [PeerJ] Analysis of synonymous codon usage patterns in sixty-four different bivalve species [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2015/1520/1/fig-3-full.png)